Link to text of accepted study (Gastroenterology, ahead of publication): Prevalence and Clinical Features of Inflammatory Bowel Diseases Associated with Monogenic Variants, Identified by Whole-exome Sequencing in 1000 Children at a Single Center
Abstract:
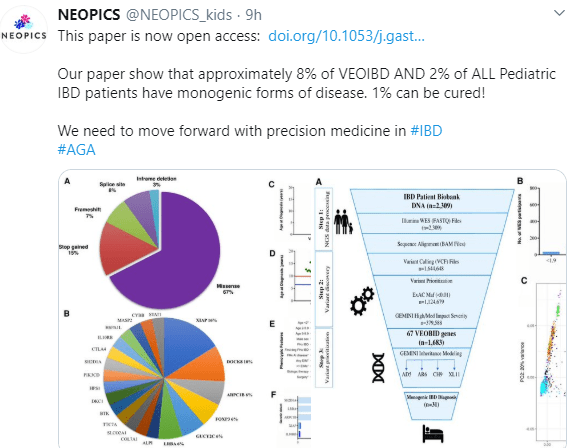
Background & Aims: A proportion of infants and young children with inflammatory bowel diseases (IBD) have subtypes associated with a single gene variant (monogenic IBD). We aimed to determine the prevalence of monogenic disease in a cohort of pediatric patients with IBD.
Methods: We performed whole-exome sequencing analyses of blood samples from an
unselected cohort of 1005 children with IBD, 0–18 y old (median age at diagnosis, 11.96 y) at a single center in Canada and their family members (2305 samples total). Variants believed to cause IBD were validated using Sanger sequencing. Biopsies from patients were analyzed by immunofluorescence and histochemical analyses.
Results: We identified 40 rare variants associated with 21 monogenic genes among 31 of the 1005 children with IBD (including 5 variants in XIAP, 3 in DOCK8, and 2 each in FOXP3, GUCY2C, and LRBA). These variants occurred in 7.8% of children younger than 6 y and 2.3% of children 6–18 y old. Of the 17 patients with monogenic Crohn’s disease, 35% had abdominal pain, 24% had non-bloody loose stool, 18% had vomiting, 18% had weight loss, and 5% had intermittent bloody loose stool. The 14 patients with monogenic ulcerative colitis or IBD unclassified received their diagnosis at a younger age, and their most predominant feature was bloody loose stool (78%). Features associated with monogenic IBD, compared to cases of IBD not associated with a single variant, were age of onset younger than 2 y (odds ratio [OR], 6.30; P=.020), family history of autoimmune disease (OR, 5.12; P=.002), extraintestinal manifestations (OR, 15.36; P<.0001), and surgery (OR, 3.42; P=.042). Seventeen patients had variants in genes that could be corrected with allogeneic hematopoietic stem cell transplantation.
Conclusions: In whole-exome sequencing analyses of more than 1000 children with IBD at a single center, we found that 3% had rare variants in genes previously associated with pediatric IBD. These were associated with different IBD phenotypes, and 1% of the patients had variants that could be potentially corrected with allogeneic hematopoietic stem cell transplantation. Monogenic IBD is rare but should be considered in analysis of all patients with pediatric onset of IBD.
KEY WORDS: HSCT, genetics, risk factor, prevalence